sklearn.cluster.AffinityPropagation#
- class sklearn.cluster.AffinityPropagation(*, damping=0.5, max_iter=200, convergence_iter=15, copy=True, preference=None, affinity='euclidean', verbose=False, random_state=None)[source]#
Perform Affinity Propagation Clustering of data.
Read more in the User Guide.
- Parameters:
- dampingfloat, default=0.5
Damping factor in the range
[0.5, 1.0)is the extent to which the current value is maintained relative to incoming values (weighted 1 - damping). This in order to avoid numerical oscillations when updating these values (messages).- max_iterint, default=200
Maximum number of iterations.
- convergence_iterint, default=15
Number of iterations with no change in the number of estimated clusters that stops the convergence.
- copybool, default=True
Make a copy of input data.
- preferencearray-like of shape (n_samples,) or float, default=None
Preferences for each point - points with larger values of preferences are more likely to be chosen as exemplars. The number of exemplars, ie of clusters, is influenced by the input preferences value. If the preferences are not passed as arguments, they will be set to the median of the input similarities.
- affinity{‘euclidean’, ‘precomputed’}, default=’euclidean’
Which affinity to use. At the moment ‘precomputed’ and
euclideanare supported. ‘euclidean’ uses the negative squared euclidean distance between points.- verbosebool, default=False
Whether to be verbose.
- random_stateint, RandomState instance or None, default=None
Pseudo-random number generator to control the starting state. Use an int for reproducible results across function calls. See the Glossary.
New in version 0.23: this parameter was previously hardcoded as 0.
- Attributes:
- cluster_centers_indices_ndarray of shape (n_clusters,)
Indices of cluster centers.
- cluster_centers_ndarray of shape (n_clusters, n_features)
Cluster centers (if affinity !=
precomputed).- labels_ndarray of shape (n_samples,)
Labels of each point.
- affinity_matrix_ndarray of shape (n_samples, n_samples)
Stores the affinity matrix used in
fit.- n_iter_int
Number of iterations taken to converge.
- n_features_in_int
Number of features seen during fit.
New in version 0.24.
- feature_names_in_ndarray of shape (
n_features_in_,) Names of features seen during fit. Defined only when
Xhas feature names that are all strings.New in version 1.0.
See also
AgglomerativeClusteringRecursively merges the pair of clusters that minimally increases a given linkage distance.
FeatureAgglomerationSimilar to AgglomerativeClustering, but recursively merges features instead of samples.
KMeansK-Means clustering.
MiniBatchKMeansMini-Batch K-Means clustering.
MeanShiftMean shift clustering using a flat kernel.
SpectralClusteringApply clustering to a projection of the normalized Laplacian.
Notes
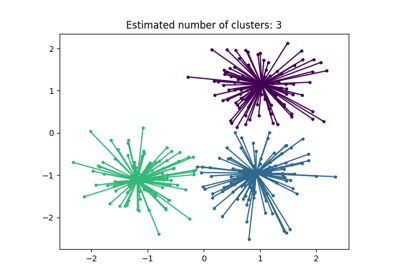
For an example, see examples/cluster/plot_affinity_propagation.py.
The algorithmic complexity of affinity propagation is quadratic in the number of points.
When the algorithm does not converge, it will still return a arrays of
cluster_center_indicesand labels if there are any exemplars/clusters, however they may be degenerate and should be used with caution.When
fitdoes not converge,cluster_centers_is still populated however it may be degenerate. In such a case, proceed with caution. Iffitdoes not converge and fails to produce anycluster_centers_thenpredictwill label every sample as-1.When all training samples have equal similarities and equal preferences, the assignment of cluster centers and labels depends on the preference. If the preference is smaller than the similarities,
fitwill result in a single cluster center and label0for every sample. Otherwise, every training sample becomes its own cluster center and is assigned a unique label.References
Brendan J. Frey and Delbert Dueck, “Clustering by Passing Messages Between Data Points”, Science Feb. 2007
Examples
>>> from sklearn.cluster import AffinityPropagation >>> import numpy as np >>> X = np.array([[1, 2], [1, 4], [1, 0], ... [4, 2], [4, 4], [4, 0]]) >>> clustering = AffinityPropagation(random_state=5).fit(X) >>> clustering AffinityPropagation(random_state=5) >>> clustering.labels_ array([0, 0, 0, 1, 1, 1]) >>> clustering.predict([[0, 0], [4, 4]]) array([0, 1]) >>> clustering.cluster_centers_ array([[1, 2], [4, 2]])
Methods
fit(X[, y])Fit the clustering from features, or affinity matrix.
fit_predict(X[, y])Fit clustering from features/affinity matrix; return cluster labels.
Get metadata routing of this object.
get_params([deep])Get parameters for this estimator.
predict(X)Predict the closest cluster each sample in X belongs to.
set_params(**params)Set the parameters of this estimator.
- fit(X, y=None)[source]#
Fit the clustering from features, or affinity matrix.
- Parameters:
- X{array-like, sparse matrix} of shape (n_samples, n_features), or array-like of shape (n_samples, n_samples)
Training instances to cluster, or similarities / affinities between instances if
affinity='precomputed'. If a sparse feature matrix is provided, it will be converted into a sparsecsr_matrix.- yIgnored
Not used, present here for API consistency by convention.
- Returns:
- self
Returns the instance itself.
- fit_predict(X, y=None)[source]#
Fit clustering from features/affinity matrix; return cluster labels.
- Parameters:
- X{array-like, sparse matrix} of shape (n_samples, n_features), or array-like of shape (n_samples, n_samples)
Training instances to cluster, or similarities / affinities between instances if
affinity='precomputed'. If a sparse feature matrix is provided, it will be converted into a sparsecsr_matrix.- yIgnored
Not used, present here for API consistency by convention.
- Returns:
- labelsndarray of shape (n_samples,)
Cluster labels.
- get_metadata_routing()[source]#
Get metadata routing of this object.
Please check User Guide on how the routing mechanism works.
- Returns:
- routingMetadataRequest
A
MetadataRequestencapsulating routing information.
- get_params(deep=True)[source]#
Get parameters for this estimator.
- Parameters:
- deepbool, default=True
If True, will return the parameters for this estimator and contained subobjects that are estimators.
- Returns:
- paramsdict
Parameter names mapped to their values.
- predict(X)[source]#
Predict the closest cluster each sample in X belongs to.
- Parameters:
- X{array-like, sparse matrix} of shape (n_samples, n_features)
New data to predict. If a sparse matrix is provided, it will be converted into a sparse
csr_matrix.
- Returns:
- labelsndarray of shape (n_samples,)
Cluster labels.
- set_params(**params)[source]#
Set the parameters of this estimator.
The method works on simple estimators as well as on nested objects (such as
Pipeline). The latter have parameters of the form<component>__<parameter>so that it’s possible to update each component of a nested object.- Parameters:
- **paramsdict
Estimator parameters.
- Returns:
- selfestimator instance
Estimator instance.
Examples using sklearn.cluster.AffinityPropagation#
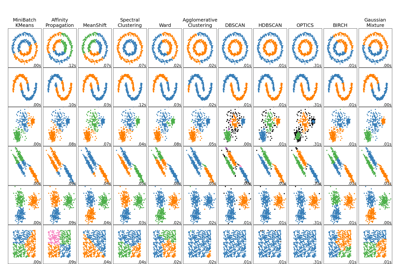
Comparing different clustering algorithms on toy datasets