sklearn.impute.KNNImputer#
- class sklearn.impute.KNNImputer(*, missing_values=nan, n_neighbors=5, weights='uniform', metric='nan_euclidean', copy=True, add_indicator=False, keep_empty_features=False)[source]#
Imputation for completing missing values using k-Nearest Neighbors.
Each sample’s missing values are imputed using the mean value from
n_neighborsnearest neighbors found in the training set. Two samples are close if the features that neither is missing are close.Read more in the User Guide.
New in version 0.22.
- Parameters:
- missing_valuesint, float, str, np.nan or None, default=np.nan
The placeholder for the missing values. All occurrences of
missing_valueswill be imputed. For pandas’ dataframes with nullable integer dtypes with missing values,missing_valuesshould be set to np.nan, sincepd.NAwill be converted to np.nan.- n_neighborsint, default=5
Number of neighboring samples to use for imputation.
- weights{‘uniform’, ‘distance’} or callable, default=’uniform’
Weight function used in prediction. Possible values:
‘uniform’ : uniform weights. All points in each neighborhood are weighted equally.
‘distance’ : weight points by the inverse of their distance. in this case, closer neighbors of a query point will have a greater influence than neighbors which are further away.
callable : a user-defined function which accepts an array of distances, and returns an array of the same shape containing the weights.
- metric{‘nan_euclidean’} or callable, default=’nan_euclidean’
Distance metric for searching neighbors. Possible values:
‘nan_euclidean’
callable : a user-defined function which conforms to the definition of
_pairwise_callable(X, Y, metric, **kwds). The function accepts two arrays, X and Y, and amissing_valueskeyword inkwdsand returns a scalar distance value.
- copybool, default=True
If True, a copy of X will be created. If False, imputation will be done in-place whenever possible.
- add_indicatorbool, default=False
If True, a
MissingIndicatortransform will stack onto the output of the imputer’s transform. This allows a predictive estimator to account for missingness despite imputation. If a feature has no missing values at fit/train time, the feature won’t appear on the missing indicator even if there are missing values at transform/test time.- keep_empty_featuresbool, default=False
If True, features that consist exclusively of missing values when
fitis called are returned in results whentransformis called. The imputed value is always0.New in version 1.2.
- Attributes:
- indicator_
MissingIndicator Indicator used to add binary indicators for missing values.
Noneif add_indicator is False.- n_features_in_int
Number of features seen during fit.
New in version 0.24.
- feature_names_in_ndarray of shape (
n_features_in_,) Names of features seen during fit. Defined only when
Xhas feature names that are all strings.New in version 1.0.
- indicator_
See also
SimpleImputerUnivariate imputer for completing missing values with simple strategies.
IterativeImputerMultivariate imputer that estimates values to impute for each feature with missing values from all the others.
References
Examples
>>> import numpy as np >>> from sklearn.impute import KNNImputer >>> X = [[1, 2, np.nan], [3, 4, 3], [np.nan, 6, 5], [8, 8, 7]] >>> imputer = KNNImputer(n_neighbors=2) >>> imputer.fit_transform(X) array([[1. , 2. , 4. ], [3. , 4. , 3. ], [5.5, 6. , 5. ], [8. , 8. , 7. ]])
For a more detailed example see Imputing missing values before building an estimator.
Methods
fit(X[, y])Fit the imputer on X.
fit_transform(X[, y])Fit to data, then transform it.
get_feature_names_out([input_features])Get output feature names for transformation.
Get metadata routing of this object.
get_params([deep])Get parameters for this estimator.
set_output(*[, transform])Set output container.
set_params(**params)Set the parameters of this estimator.
transform(X)Impute all missing values in X.
- fit(X, y=None)[source]#
Fit the imputer on X.
- Parameters:
- Xarray-like shape of (n_samples, n_features)
Input data, where
n_samplesis the number of samples andn_featuresis the number of features.- yIgnored
Not used, present here for API consistency by convention.
- Returns:
- selfobject
The fitted
KNNImputerclass instance.
- fit_transform(X, y=None, **fit_params)[source]#
Fit to data, then transform it.
Fits transformer to
Xandywith optional parametersfit_paramsand returns a transformed version ofX.- Parameters:
- Xarray-like of shape (n_samples, n_features)
Input samples.
- yarray-like of shape (n_samples,) or (n_samples, n_outputs), default=None
Target values (None for unsupervised transformations).
- **fit_paramsdict
Additional fit parameters.
- Returns:
- X_newndarray array of shape (n_samples, n_features_new)
Transformed array.
- get_feature_names_out(input_features=None)[source]#
Get output feature names for transformation.
- Parameters:
- input_featuresarray-like of str or None, default=None
Input features.
If
input_featuresisNone, thenfeature_names_in_is used as feature names in. Iffeature_names_in_is not defined, then the following input feature names are generated:["x0", "x1", ..., "x(n_features_in_ - 1)"].If
input_featuresis an array-like, theninput_featuresmust matchfeature_names_in_iffeature_names_in_is defined.
- Returns:
- feature_names_outndarray of str objects
Transformed feature names.
- get_metadata_routing()[source]#
Get metadata routing of this object.
Please check User Guide on how the routing mechanism works.
- Returns:
- routingMetadataRequest
A
MetadataRequestencapsulating routing information.
- get_params(deep=True)[source]#
Get parameters for this estimator.
- Parameters:
- deepbool, default=True
If True, will return the parameters for this estimator and contained subobjects that are estimators.
- Returns:
- paramsdict
Parameter names mapped to their values.
- set_output(*, transform=None)[source]#
Set output container.
See Introducing the set_output API for an example on how to use the API.
- Parameters:
- transform{“default”, “pandas”}, default=None
Configure output of
transformandfit_transform."default": Default output format of a transformer"pandas": DataFrame output"polars": Polars outputNone: Transform configuration is unchanged
New in version 1.4:
"polars"option was added.
- Returns:
- selfestimator instance
Estimator instance.
- set_params(**params)[source]#
Set the parameters of this estimator.
The method works on simple estimators as well as on nested objects (such as
Pipeline). The latter have parameters of the form<component>__<parameter>so that it’s possible to update each component of a nested object.- Parameters:
- **paramsdict
Estimator parameters.
- Returns:
- selfestimator instance
Estimator instance.
- transform(X)[source]#
Impute all missing values in X.
- Parameters:
- Xarray-like of shape (n_samples, n_features)
The input data to complete.
- Returns:
- Xarray-like of shape (n_samples, n_output_features)
The imputed dataset.
n_output_featuresis the number of features that is not always missing duringfit.
Examples using sklearn.impute.KNNImputer#
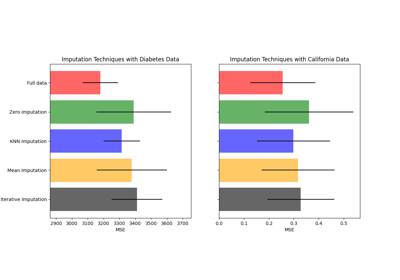
Imputing missing values before building an estimator